Pfizer/BioNTech C4591001 Trial - Safety Population, with & without prior infection
How to magically alter your safety population with a cheap database trick
Introduction
This article is the product of a joint effort with Josh Guetzkow1.
We detail here a new set of anomalies affecting the safety analysis.
The subject’s “eligibility to the safety population” - the population used to determine if the product was safe - was determined in code by a set of variables.
Failure to set one of these variable properly has allowed the sponsors to disregard a considerable quantity of subjects, disproportionately affecting the treatment group, and illustrates that the code provided, and the data provided, don’t correspond.
The scripts provided below assume that you have setup your project as described in this previous article2.
Safety population at baseline
Two pages on the Month 6 interim report3 feature AEs based on the “Covid at baseline” status of the subjects, which are pasted below (on the left the 1379 subjects who had COVID at baseline - on the right the 42 194 subjects who didn’t.


Manually reporting these figures in a table, the populations featured in these two tables, for the two treatment arms, can be summarized as follows:
The total population in this safety analysis is therefore of 43 573 subjects.
This figure is surprising, as “43 573” is detailed absolutely nowhere in the BLA memorandum4 populations breakdown, or the corresponding study, by Stephen J. Thomas et al. (published on MedRxiv5 on July 28, 2021 and in the NEJM6 on November 4, 2021).
The Analyst Data Reviewer Guide (ADRG)7 provided by Pfizer to the FDA defines the safety population as follows, page 81, and reports 43 847 subjects.
We verified the population documented in the ADRG, by applying the filters documented directly in the ADSL file, via a first dedicated R script.
It indeed results in 43 847 subjects.
Reaching the 43 573 figure is a bit more tricky as it is “documented” very discretely in the ADRG.
Page 12 states that “ADSL.COVBLST” - the tag in the subject level data set to “POS” indicating if the subject had Covid or not at dose 1… has two possible statuses (Positive or Negative)…
… But to be fair, page 64 does mention that this status could be missing.
There are 274 subjects without “COVBLST” value, in this 43 847 subjects “Safety population”.
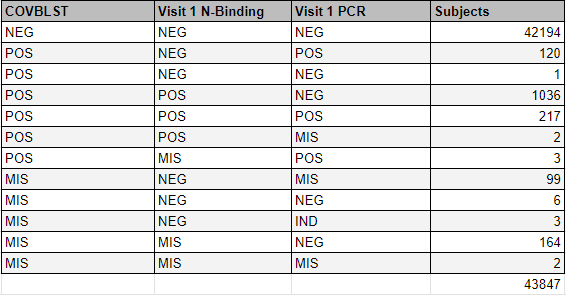
Values distribution for this COVBLST tag, in the 43 847 safety population, are represented in this table.
The impact of this simple tag being “missing” was that these 274 subjects weren’t included in these specific “COVID at baseline” analyses.
Reviewing the code populating the data
We have to identify precisely how the variable was set, to refer to the SAS code provided8.
We don’t have the raw data, so we can’t run it, but we can investigate how it worked.
To simplify:
The code first gathered and processed various data, including body mass index (BMI), comorbidities, and COVID-19 related information.
It then checked for the presence of COVID-19 by looking at different factors. These include medical history (MHDECOD) related to COVID-19, a positive result for a SARS-CoV-2 test (mborres="POS" and mbdy<=1), and a positive result for a N-binding antibody test (isorres="POS" and isdy<=1).
If any of these conditions were met, the individual was marked as "POS" for COVBLST, indicating they had COVID-19 at the baseline of the study.
If none of these conditions were met, and the individual had a negative result for both the SARS-CoV-2 test and the N-binding antibody test, they were marked as "NEG" for COVBLST, indicating they did not have COVID-19 at the baseline of the study.
The presence of subjects with neither "POS" nor "NEG" in the COVBLST tag could therefore be due to a few reasons:
Missing Data: The tests for SARS-CoV-2 or N-binding antibody might not have been conducted for these subjects, or the results of these tests might not have been recorded properly. This could result in missing data for these subjects.
Timing of Tests: The COVBLST tag is determined on Positive tests based on tests conducted on or before the first day of the study (mbdy<=1 and isdy<=1). If the tests were conducted after the first day, the results would not be considered in determining the COVBLST tag and essentially treated as missing.
Medical History: The COVBLST tag also considers the medical history of the subjects. If the subjects do not have a medical history related to COVID-19, and if they have not been tested for SARS-CoV-2 or N-binding antibody, they would not be categorized as either "POS" or "NEG".
In absence of the raw data, our first step is to review the status of the subjects, and their time between V1 tests & Dose 1, to check if point 2 above can explain the anomaly of 274 subjects missing COVBLST.
COVBLST status, V1 N-Binding & PCR Status
The distribution of the possible values for V1 N-Binding, PCR, and COVBLST status, is represented in the table below. The visit 1 tests are merged and normalized using this second script; and the distribution on the safety population, along with the subsequent analysis, are generated by this third.
This distribution raises more questions than it answers. If negative tests present for both N-binding or PCR were sufficient for the tag to be set, why would we have tags not set for 6 subjects, while both results are known, and negative ?
COVBLST status, V1 N-Binding & PCR Status, Days from Dose to Tests
We generated another distribution, including the days of differences between the day of the tests, and the day of the first dose.
We can observe that subjects with 18 days, or more, between their visit 1 tests & their dose 1 all have “Negative” value for COVBLST.
We can only wonder how it could happen for subjects to have 18 days (or more) between dose 1 & the very testing supposed to ensure if the subject had contracted COVID at dose 1.
Arms impacted by this anomaly
Lastly, we’re left with the task to review which treatment arms were impacted by this anomaly.
An important note is that subjects were “unblinded” very early in the trial, as extensively detailed in this article9.
Therefore a large amount of subjects who received Placebo also received the active product (“cross-over subjects”).
The subjects by arms, depending if their tag is set or not set, are further synthesized in the tables below.
We therefore have:
21 772 + 19 414 = 41 186 subjects who received BNT162b2 at some stage of the trial, and have their “COVBLST” tag set.
261 subjects (0,63%) who received BNT162b2 at some stage of the trial, and haven’t had their “COVBLST” tag set.
2 387 subjects who received only Placebo and have their “COVBLST” tag set.
13 subjects (0,54%) who only received Placebo, and haven’t had their “COVBLST” tag set.
We have an imbalance which has impacted more the BNT162b2 group, but it fails to achieve statistical significance.
Sites impacted by this anomaly
A breakdown of the anomaly by sites shows further details, highlighting as often abnormal behaviors at sites scale.
A handful of sites have been affected on more than 10% of their subjects in a given treatment arm - with up to 19% of their BNT162b2 concerned on site 1169.
Cover picture credits to
(or it might have been taken in the FDA’s entry hall - sources diverge).phmpt.org/wp-content/uploads/2022/03/FDA-CBER-2021-5683-0022867-to-0023006_125742_S1_M5_c4591001-A-P-adsl-sas.txt, line 2067 to 2562, * Specification 4 *